
水合三环己基锡茶碱-7-乙酸配合物的合成、结构及性能
-
关键词:
- 水合三环己基锡茶碱-7-乙酸配合物
- / 合成
- / 晶体结构
- / 体外抗肿瘤活性
English
Synthesis, structure, and properties of hydrated tricyclohexyltin theophylline-7-acetic acid complex
-
0. Introduction
Organotin complexes have attracted considerable attention due to their versatile structures, rich reactivity, notable biological activity, and broad applications[1-8]. Notably, their inhibitory effects on cancer cell proliferation have opened new avenues for developing broad-spectrum and efficient antitumor drugs[9-15]. The structural, reactive, and biological properties of organotin complexes are influenced not only by the hydrocarbyl groups directly bonded to the tin atom but also by properties of the ligands. Functionalized ligands can significantly alter the coordination mode of tin atoms, thereby modulating the balance between toxicity and biological activity[16-22]. Especially when substituted carboxylic acids or heterocyclic carboxylic acids containing active groups as substituents are used as ligands, they are more likely to form organic tin complexes with distinct structural features and unique properties[23-28]. Theophylline-7-acetic acid is a prime example of this type of ligand.
Theophylline-7-acetic acid (tpH), a xanthine derivative, exhibits diverse biological activities, including adenosine receptor antagonism, protein arginine deiminase activation, bronchodilation, and inhibition of cAMP phosphodiesterase isoenzymes in rat lung. It also shows potential in treating adipose or "orange peel" skin. Studies have demonstrated its binding affinity for the A2B adenosine receptor in HEK-293 human stem cells. As a multidentate ligand, tpH can coordinate with metal ions in various modes (monodentate, bidentate, or multidentate), forming complexes with novel structures stabilized by hydrogen bonds and π-π stacking interactions[29-30].
In this study, we synthesized the hydrated tricyclohexyltin tpH complex [Sn(C6H11)3(tp)(H2O)] and characterized it using IR, NMR, elemental analysis, and powder X-ray diffraction (PXRD). The crystal structure was elucidated via single-crystal X-ray diffraction, and quantum chemistry calculations were employed to explore its stability and electronic properties. Furthermore, the complex′s thermal stability, electrochemical properties, and in vitro anticancer activity were investigated.
1. Experimental
1.1 Instruments and reagents
Infrared spectra were recorded on a Shimadzu FTIR-8700 spectrometer (Shimadzu, Japan) using KBr pellets (400-4 000 cm-1). Elemental analysis was performed with a PE-2400 Elemental Analyzer (PerkinElmer Inc., USA). 1H and 13C NMR spectra were obtained using a Bruker Avance Ⅲ HD 500 MHz instrument (Bruker, Switzerland). The crystal structure was determined using a Bruker Smart Apex Ⅱ CCD single-crystal diffractometer (Bruker, Germany). Melting points were measured with an X-4 micro-digital melting point apparatus (Beijing Tech Instrument Company, China). Electrochemical studies were conducted on a CHI660D electrochemical workstation (Shanghai Chenhua, China), and thermogravimetric analysis (TGA) was performed using a Thermal Analyst TG209F3 instrument (Netzsch, Germany). The PXRD data were measured using an XRD-6100 X-ray diffractometer (Shimadzu, Japan) under the following conditions: working voltage of 40.0 kV, current of 30.0 mA, Cu Kα radiation, wavelength of 0.154 06 nm, and a scanning range of 5°-80°. All reagents are analytical grade.
1.2 Synthesis
A mixture of absolute ethanol (30 mL), tricyclohexyltin hydroxide (0.385 g, 1 mmol), and tpH (0.237 g, 1 mmol) was sealed in a 50 mL pressure-resistant bottle and stirred at 110 ℃ for 3 h. After some solvents were removed by rotary evaporation, the mixture was allowed to stand to precipitate a white solid. The solid was recrystallized from an appropriate solvent to obtain 0.476 g of colorless crystals. Yield: 76.40%. m.p. 114-116 ℃. Anal. Calcd. for C27H44N4O5Sn(%): C, 52.01; N, 8.99; H, 7.13. Found(%): C, 52.23; N, 8.96; H, 7.09. IR data(KBr, cm-1): ν(O—H) 3 429; ν(C—H) 3 108, 2 920, 2 846; ν(C=O) 1 704; νas(COO) 1 664; νs(COO) 1 348; ν(Sn—C) 590; ν(Sn—O) 506, 442. 1H NMR (500 MHz, CDCl3): δ 7.58 (s, 1H), 5.02 (s, 2H), 3.58 (s, 3H), 3.36 (s, 3H), 1.93-1.87 (m, 9H), 1.63-1.59 (m, 15H), 1.32-1.27 (m, 9H). 13C NMR (126 MHz, CDCl3): δ 170.90, 155.08, 151.78, 148.32, 141.72, 107.39, 48.27, 34.34, 30.91, 29.71, 28.79, 27.74, 26.75.
1.3 Crystal structure determination
A single crystal with a size of 0.21 mm×0.19 mm×0.18 mm was selected, and the diffraction experiment was carried out on the Bruker SMART APEX Ⅱ single crystal diffractometer. All data were collected at 296(2) K by φ-ω scanning using graphite monochromatic Mo Kα radiation (λ=0.071 073 nm). The diffraction intensity data were corrected by multiple scanning absorption. Most of the non-hydrogen atoms in the crystal structure were solved by the direct method, and the rest of the non-hydrogen atoms were located by difference Fourier syntheses. The position coordinates of the hydrogen atoms in the unit cell were given by the theoretical evaluation of the hydrogenation reactions. The coordinates of all non-hydrogen atoms and anisotropic temperature factors were refined by full-matrix least-squares. All structural analysis was completed by calling the SHELX-97 program on WINGX. Selected crystallographic data of the complex are given in Table 1.
Table 1
Parameter [Sn(C6H11)3(tp)(H2O)] Empirical formula C27H44N4O5Sn Formula weight 623.35 Crystal system Orthorhombic Space group Iba2 a / nm 1.954 11(10) b / nm 1.702 11(9) c / nm 1.754 22(9) V / nm3 5.834 7(5) Z 8 Dc / (Mg·m-3) 1.419 μ(Mo Kα) / cm-1 9.17 F(000) 2 592 θ range for data collection / (°) 1.587-29.090 Index range -19 ≤ h ≤ 26,
-23 ≤ k ≤ 22,
-22 ≤ l ≤ 24Reflection collected 18 987 Reflection collected, unique 7 010 (Rint=0.033 2) Goodness-of-fit on F 2 1.025 Final R indices R1, wR2 [I > 2σ(I)] 0.031 9, 0.063 6 R indices R1, wR2 (all data) 0.049 1, 0.071 1 Largest diff. peak and hole / (e·nm-3) 614 and -346 1.4 In vitro anticancer activity of the complex
The growth inhibitory effect toward tumor cells was evaluated by means of the CCK-8 assay. Briefly, 3×103-8×103 cells per well, dependent upon the growth characteristics of the cell line, were seeded in 96-well microplates in growth medium (100 μL) with a discharge gun. After 24 h, the medium was removed, and the complex (10 mmol·L-1) was added to the plate with a gradient dilution of the culture medium (maximum working concentration 10 μmol·L-1, dilution ratio 1∶3). Triplicate cultures were established for each treatment. After 72 h, each well was treated with 10 μL CCK-8 solution and incubated for 1-4 h. Cell growth inhibition was detected by measuring the absorbance of each well at 450 and 650 nm using a super microplate reader. The absorbance value was standardized in the control wells, and the cell survival rate was calculated. The inhibitory concentration at 50% of the inhibitory rate (IC50) was calculated using GraphPadPrism 8.0 Software.
2. Results and discussion
2.1 Spectroscopic properties of the complex
In the IR spectrum, the absorption peak of the complex at 590 cm-1 is the Sn—C bond absorption peak. The absorption peaks appearing at 506 and 442 cm-1 are the absorption peaks of Sn—O bonds formed by water molecule oxygen and carboxyl oxygen with tin, suggesting the presence of two different Sn—O bonds in the complex molecule. The asymmetric stretching vibration peak of the carboxyl group [νas(COO)] in the complex was at 1 664 cm-1, and the symmetric stretching vibration absorption peak [νs(COO)] was at 1 348 cm-1. Its Δν [the difference between νas(COO) and νs(COO)] was 316 cm-1, which was greater than 200 cm-1, indicating that the carboxyl group in the complex coordinates with tin as monodentate oxygen. The complex exhibited a broad absorption peak at 3 429 cm-1, which is an O—H bond absorption peak, showing the presence of water molecules in the crystal.
In the 1H NMR spectrum, the single peak appearing at δ 7.58 is the absorption peak of H on the imidazole ring carbon atom, with a total of one proton. A single peak appeared at δ 5.02, which is the proton absorption peak on the acetic acid methyl carbon, with a total of two protons. The spectrum exhibits a single peak at δ 3.58 and 3.02, representing the absorption peaks of methyl H on the nitrogen atom of the pyrimidine ring, with three protons each. There were multiple peaks between δ 1.93 and 1.27, which are the absorption peaks of protons on cyclohexyl, with a total of 33 protons. The carboxyl carbon absorption peak (δ 170.90), the aromatic carbon absorption peak on the purine ring (δ 155.08, 151.78, 148.32, 141.72, 107.39), and the absorption peaks of saturated carbon (δ 48.27, 34.34, 30.91, 29.71, 28.79, 27.74, 26.75) appeared in the 13C NMR spectrum.
The results obtained by IR and NMR are consistent with the crystal structure determined by single-crystal X-ray diffraction.
2.2 Crystal structure description
The main bond lengths and bond angles of the complex are listed in Table 2, and the molecular structure is shown in Fig.1.
Table 2
Sn1—O3 0.213 1(3) Sn—C16 0.216 0(4) Sn1—O5 0.259 8(3) Sn1—C2 0.215 8(5) Sn1—C1 0.217 2(6) O3—Sn1—C2 91.72(19) C22—Sn1—C10 113.55(18) C22—Sn1—O5 82.99(17) O3—Sn1—C16 100.04(14) C16—Sn1—C10 126.8(2) C16—Sn1—O5 79.16(14) C22—Sn1—C16 114.83(19) O3—Sn1—O5 173.65(14) C10—Sn1—O5 86.15(17) O3—Sn1—C10 99.2(2) Figure 1
Based on the data presented in Fig. 1 and the structural parameters, the complex adopts a mononuclear structure. The central tin atom coordinates with three cyclohexyl carbon atoms, one carboxylate oxygen atom, and one water oxygen atom, forming a five-coordinated, severely distorted trigonal bipyramidal geometry. In this structure, three cyclohexyl carbon atoms occupy equatorial positions, defining the equatorial plane. The carboxylate oxygen atom (O3) and the water oxygen atom (O5) occupy the two axial positions above and below this plane. The sum of the bond angles between the three equatorial carbon atoms (C22—Sn1—C10, C10—Sn1—C16, C16—Sn1—C22) is 355.18°, deviating significantly from the ideal 360°. This indicates that these three atoms are not strictly coplanar. The bond angles between the axial carboxylate oxygen atom (O3) and the equatorial carbon atoms are 91.72° (O3—Sn1—C22), 99.2° (O3—Sn1—C10), and 100.04° (O3—Sn1—C16), all greater than 90°. In contrast, the bond angles between the axial water oxygen atom (O5) and the equatorial carbon atoms are 86.15° (C10—Sn1—O5), 82.99° (C22—Sn1—O5), and 79.16° (C16—Sn1—O5), all less than 90°. This distortion arises from the steric effect of the theophylline-7-acetoxy group, which forces the three cyclohexyl groups towards the water molecule. Consequently, the cyclohexyl groups adopt different spatial environments, resulting in asymmetric, chair-like six-membered rings. Furthermore, the axial O3—Sn1—O5 bond angle is 173.65°, deviating significantly from the ideal linear angle of 180.0°. Thus, the tin atom exhibits a severely distorted trigonal bipyramidal coordination geometry within the molecule.
Additionally, within the crystal, the complex molecules form a 2D network structure through hydrogen bonds involving coordinated water molecules and π-π stacking interactions, as shown in Fig.2. Among these, the hydrogen bond length of H5B⋯O2 is 0.201 8(3) nm, and that of H5A⋯N2 is 0.203 6(4) nm. There are continuous and strong π-π stacking interactions between the six-membered rings on adjacent purine ring planes (Cg: C2-N3-C3-N4-C4-C5; coordinates: 0.951 83, 0.412 70, 0.913 16). The interplanar distance between adjacent planes is 0.351 8 nm, and the dihedral angle between the planes is 6.936°. Through π-π stacking and O—H⋯N hydrogen bonding, the complex forms a more stable 2D network structure.
Figure 2
2.3 Energy and frontier molecular orbital composition of the complex
According to the atomic coordinates of the crystal structure, quantum chemical calculations were performed using the Gaussian 03W program at the B3LYP/LANL2DZ level. The calculated total energy (ET) of the molecule is -7 582.567 897 6 a.u. The energy of the highest occupied molecular orbital (EHOMO) is -0.222 71 a.u., and the energy of the lowest unoccupied molecular orbital (ELUMO) is 0.236 16 a.u. Both the total energy and the HOMO energy are relatively low. The HOMO-LUMO energy gap (ΔE) is calculated to be 0.458 87 a.u., indicating a stable molecular structure. Furthermore, from the perspective of redox processes, the low-lying HOMO energy level suggests that the molecule is difficult to oxidize, as it is reluctant to donate electrons.
To explore the electronic structure and bonding characteristics of the complex, molecular orbital (MO) analysis was performed. The contribution of a specific part to a molecular orbital was represented by the sum of the squares of the coefficients of the participating atomic orbitals, subsequently normalized. The atoms of the complex were partitioned into eight distinct groups: (a) tin atom (Sn); (b) carboxylate oxygen atom (O1); (c) carboxylate carbon atom (C1); (d) non-hydrogen atoms of the ligand excluding the carboxylate group (C, N, O), denoted as L; (e) cyclohexyl carbon atom C2 bonded to the tin atom; (f) other cyclohexyl carbon atoms (C3); (g) oxygen atom O2 of the coordinated water molecule; (h) hydrogen atoms (H). The five highest-energy occupied molecular orbitals and the five lowest-energy unoccupied molecular orbitals were selected for detailed analysis. The computational results are summarized in Table 3 and illustrated in Fig.3.
Table 3
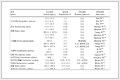 Table 3. Calculated some frontier molecular orbitals composition of the complex at the B3LYP/LANL2DZ level
Table 3. Calculated some frontier molecular orbitals composition of the complex at the B3LYP/LANL2DZ levelε / a.u. Sn O1 C1 L C2 C3 O2 H 158 -0.298 93 18.004 16 23.815 09 0.416 21 1.880 56 31.111 49 18.967 60 0.609 72 5.607 34 159 -0.282 94 1.727 00 41.748 25 1.286 45 52.236 33 1.529 51 0.530 76 0.409 20 1.824 77 160 -0.274 13 0.183 39 6.210 45 0.274 62 91.249 12 0.314 41 0.134 85 0.029 10 1.863 13 161 -0.271 55 1.567 41 82.126 74 2.184 50 4.520 87 7.452 52 3.317 19 0.005 92 1.004 55 162HO -0.222 71 4.157 04 1.460 73 0.388 42 89.181 90 3.090 14 0.051 13 0.020 44 2.030 33 163LU 0.236 16 0.153 17 1.646 15 1.079 03 91.609 41 0.108 92 0.018 57 0.005 94 0.649 58 164 0.321 81 53.155 39 2.408 11 0.284 80 0.719 14 28.141 96 5.905 84 6.643 74 3.014 85 165 0.330 03 0.299 42 0.089 97 0.165 88 98.279 37 0.265 69 0.099 08 0.014 06 0.948 18 166 0.344 15 0.472 36 0.196 20 0.908 53 98.452 24 0.284 39 0.051 23 0.000 68 0.535 92 167 0.359 22 3.120 34 40.046 67 47.584 89 48.463 97 2.425 80 0.417 19 0.007 57 5.516 55 Figure 3
In the composition of the HOMO, the non-hydrogen atoms of the ligand, excluding the carboxylate group, make the largest contribution, accounting for 89.18%. This is followed by the tin atom, the cyclohexyl carbon atom bonded to tin, and the carboxylate oxygen atom, contributing 4.16%, 3.09%, and 1.46%, respectively. Contributions from other atoms are relatively minor; notably, the oxygen atom of the coordinated water molecule contributes only 0.02%. These results indicate that the theophylline moiety of the ligand as a whole contributes significantly to the stability of the molecule, and the Sn—C and Sn—O (carboxylate O1) bonds in the structure are relatively stable, signifying strong bonding interactions between the tin atom and both the cyclohexyl carbon atom and the carboxylate oxygen atom. In contrast, the bonding interaction with the coordinated water molecule is comparatively weak.
Comparison of the orbital compositions of the HOMO and the LUMO reveals that upon electronic excitation from the ground state to the excited state, charge transfer occurs primarily from the cyclohexyl group atoms and the tin atom to the orbitals localized on the ligand.
2.4 Thermal stability
The TG curve of the complex was recorded using a TG209F3 thermogravimetric analyzer under an air atmosphere. As shown in Fig. 4, the weight loss of the complex occurred primarily in two distinct stages. In the first stage, a gradual weight loss commenced at 110 ℃ and essentially ceased by 140 ℃, resulting in a total weight loss of 3.21%. This stage is attributed to the loss of coordinated water molecules, and the observed value is in good agreement with the calculated value of 2.89%. The second stage involved the decomposition of the organic ligands: weight loss initiated at 240 ℃, progressively accelerated, then decelerated around 395 ℃, and finally stabilized near 575 ℃. The total weight loss in this stage was 74.25%, corresponding primarily to the decomposition of the tp- ligand and the cyclohexyl groups. The final residue weight stabilized at 22.54%. This residue is tentatively assigned as SnO2, which also shows reasonable agreement with the calculated value of 24.19%.
Figure 4
2.5 Electrochemical property
Cyclic voltammetry (CV) of the complex was performed on a CHI660D electrochemical workstation (Shanghai Chenhua) using a three-electrode system: a glassy carbon working electrode, a saturated calomel reference electrode, and a platinum auxiliary electrode. The electrolyte was a mixture of 5 mL ethanol and 5 mL acetate buffer solution (1.65 mol·L-1). Scanning rates ranged from 0.01 to 0.20 V·s-1.
Fig. 5A shows the influence of different scan rates on the electrochemical behavior of the complex. It is seen that all curves exhibited distinct reduction peaks, with no distinct oxidation peaks appearing. This indicates that the complex undergoes a completely irreversible electrochemical reduction process. When the scan rate increased from 0.01 to 0.20 V·s-1, as seen in the figure, the peak potential gradually changed, and the reduction peak potential shifted toward the negative direction. According to the relation between peak potential (Ep) and the logarithm of scan rate (ln v) (Fig. 5B), it could be calculated that the complex undergoes an electrochemical reduction process involving one electron and one proton. This may correspond to the carbonyl group on the pyrimidine ring of the ligand obtaining a proton and becoming a hydroxyl group. The electrochemical analysis results reflect that the ligand can still maintain its electroreduction performance after coordinating with the metal center Sn.
Figure 5
2.6 PXRD analysis
To verify the purity of the complex, a PXRD test was performed. Fig. 6 presents the simulated PXRD pattern alongside the experimental PXRD pattern. Correspondence in position and intensity was observed for most diffraction peaks between the simulated and experimental patterns, indicating high crystalline purity of the synthesized complex.
Figure 6
2.7 In vitro antitumor activity
The in vitro growth inhibitory activity of the complex against human gastric adenocarcinoma cells (AGS), human acute lymphoblastic leukemia cells (MOLT-4), and human breast cancer cells (MDA-MB-231) was tested and compared with paclitaxel as a control, as shown in Table 4. The results indicate that the inhibitory activity of the complex against the studied cancer cells was stronger than that of the bis(o-fluorobenzyl)tin 2,2′-bipyridine-6,6′-dicarboxylate complex and the di-n-butyltin 2,2′-bipyridine-6,6′-dicarboxylate complex reported previously[28]. This reflects the influence of different structures of the alkyl group and ligand on the anticancer activity of the organotin complex. However, compared to paclitaxel, the complex′s activity against MOLT-4 cells was weaker, as was its activity against the other two cell lines. The detailed biological activity of the complex requires further in-depth investigation.
Table 4
Compound IC50 / (μmol·L-1) AGS MOLT4 MDA-MB-321 [Sn(C6H11)3(tp)(H2O)] 2.6 1.433 7.061 Paclitaxel 0.894 4.587 0.004 3. Conclusions
The hydrated tricyclohexyltin theophylline-7-acetic acid complex [Sn(C6H11)3(tp)(H2O)] was synthesized via an ethanol solvothermal method. In vitro antitumor activity tests indicated that the complex exhibited significant inhibitory effects against human gastric adenocarcinoma cells (AGS), human acute lymphoblastic leukemia cells (MOLT4), and human breast cancer cells (MDA-MB-231).
-
-
[1]
YADAV V K, NATH M. Diorganotin (Ⅳ) derivatives of quinoline-2-carboxaldehyde 4-phenylthio-semicarbazone: Synthesis, spectroscopic characterization, the crystal structure of [Bu2Sn(QCP)Cl], DFT studies, CT-DNA binding, and plasmid-DNA cleavage studies[J]. J. Mol. Struct., 2025, 1319: 139594 doi: 10.1016/j.molstruc.2024.139594
-
[2]
PANDEY P, WALAWALKAR M G, MURUGAVEL R. Luminescent 8-hydroxyquinoline derived tin(Ⅳ) complexes[J]. J. Org. Chem., 2024, 1007: 123023 doi: 10.1016/j.jorganchem.2024.123023
-
[3]
TAN Y X, QIN Q Q, LI A D, FU W W, JIANG W J. Synthesis, characterization, and biological activity of six di-p-chlorobenzyltin complexes derived from ONO tridentate ligands[J]. J. Org. Chem., 2023, 1002: 122905 doi: 10.1016/j.jorganchem.2023.122905
-
[4]
PARVEEN B, SHAHZADI S, ALI S, FEIZI-DEHNAYEBI M, MUNAWAR K S, ASHFAQ M, TAHIR M N. Synthesis, spectral characterizations, computational studies and biological investigation of 4-(4-(2-hydroxyethyl)phenylamino)-4-oxobutanoic acid and its trimethyltin(Ⅳ) complex[J]. J. Mol. Struct., 2024, 1315: 138851 doi: 10.1016/j.molstruc.2024.138851
-
[5]
SU H Q, ZHANG R F, GUO Q, WANG J, LI Q L, DU X M, RU J, ZHANG Q F, MA C L. Five organotin complexes derived from hydroxycinnamic acid ligands: Synthesis, structure, in vitro cytostatic activity and binding interaction with BSA[J]. J. Mol. Struct., 2022, 247: 131290-131301
-
[6]
RUAN B F, TIAN Y P, ZHOU H P, WU J Y, HU R T, ZHU C H, YANG J X, ZHU H. Synthesis, characterization and in vitro antitumor activity of three organotin(Ⅳ) complexes with carbazole ligand[J]. Inorg. Chim. Acta, 2011, 365(1): 302-308 doi: 10.1016/j.ica.2010.09.024
-
[7]
冯泳兰, 邝代治, 张复兴, 庾江喜, 蒋伍玖, 朱小明. 两个具有Sn4O4梯状结构二丁基锡羧酸酯的微波溶剂热合成、结构和体外抗癌活性[J]. 无机化学学报, 2017, 33(5): 830-836FENG Y L, KUANG D Z, ZHANG F X, YU J X, JIANG W J, ZHU X M. Two di-n-butyltin carboxylates with a Sn4O4ladder-like framework: Microwave solvothermal syntheses, structures and in vitro antitumor activities[J]. Chinese J. Inorg. Chem., 2017, 33(5): 830-836
-
[8]
邝代治, 庾江喜, 冯泳兰, 朱小明, 蒋伍玖, 张复兴. 大环超分子二(三环己基锡)吡啶-二甲酸酯的合成、结构和抗癌活性[J]. 无机化学学报, 2018, 34(6): 1035-1042KUANG D Z, YU J X, FENG Y L, ZHU X M, JIANG W J, ZHANG F X. Syntheses, structures and in vitro antitumor activity of bis(tricyclohexyltin) pyridinedicarboxylate with macrocyclic supramolecular structure[J]. Chinese J. Inorg. Chem., 2018, 34(6): 1035-1042
-
[9]
JIANG W J, FAN S J, ZHU Z H, HUANG H F, TAN Y X, ENG Y Y. Design, synthesis and mechanistic studies of novel arylformylhydrazone butylphenyltin complexes as potential anticancer agents[J]. Bioorg. Chem., 2024, 149: 107502 doi: 10.1016/j.bioorg.2024.107502
-
[10]
TAN Y X, ZHANG Z J, LIU J Z, TAN Y J, JIANG W J. Syntheses, crystal structures, and anticancer activities of organotin carboxylates based on Alrestatin[J]. J. Mol. Struct., 2025, 1322: 140697 doi: 10.1016/j.molstruc.2024.140697
-
[11]
卿菁菁, 何帆, 刘智辉, 侯帅鹏, 刘娅, 蒋一凡, 谭梦婷, 何丽芳, 张复兴, 朱小明. 两个丁二肟有机锡配合物的合成、结构及抗癌活性[J]. 无机化学学报, 2024, 40(7): 1301-1308QING Q Q, HE F, LIU Z H, HOU S P, LIU Y, JIANG Y F, TAN M T, HE L F, ZHANG F X, ZHU X M. Synthesis, structure, and anticancer activity of two complexes of dimethylglyoximeorganotin[J]. Chinese J. Inorg. Chem., 2024, 40(7): 1301-1308
-
[12]
VIEIRA F T, LIMA G M, MAIA J R S, SPEZIALI N L, ARDISSON J D, RODRIGUES L, JUNIOR A C, ROMERO O B. Synthesis, characterization and biocidal activity of new organotin complexes of 2-(3-oxocyclohex-1-enyl)benzoic acid[J]. Eur. J. Med. Chem., 2010, 45: 883-889 doi: 10.1016/j.ejmech.2009.11.026
-
[13]
XIAO X, LIANG J W, XIE J Y, LIU X, ZHU D S, DONG Y. Organotin(Ⅳ) carboxylates based on 2-(1, 3-dioxo-1H-benzo[de]-isoquinolin-2(3H)-yl)acetic acid: Syntheses, crystal structures, luminescent properties and antitumor activities[J]. J. Mol. Struct., 2017, 1146: 233-241 doi: 10.1016/j.molstruc.2017.05.141
-
[14]
YUSOF E N M, LATIF M A M, TAHIR, M I M, SAKOFF J A, VEERAKUMARASIVAM A, PAGE A J, TIEKINK E R T, RAVOOF T B S A. Homoleptic tin(Ⅳ) compounds containing tridentate ONS dithiocarbazate Schiff bases: Synthesis, X-ray crystallography, DFT and cytotoxicity studies[J]. J. Mol. Struct., 2020, 1205: 127635-127643 doi: 10.1016/j.molstruc.2019.127635
-
[15]
邓欣, 张复兴, 卿菁菁, 杨舸, 何帆, 何丽芳, 刘智辉, 侯帅鹏. 两个苯甲羟肟酸有机锡配合物的合成、结构及抗癌活性[J]. 无机化学学报, 2023, 39(11): 2083-2090DENG X, ZHANG F X, QING J J, YANG G, HE F, HE L F, LIU Z H, HOU S P. Synthesis, structure and anticancer activity of two benzohydroxamic acid organotin complexes[J]. Chinese J. Inorg. Chem., 2023, 39(11): 2083-2090
-
[16]
TAN Y X, HUANG H F, ZANG C W, YUAN S T, JIANG W J. Molecular structure regulation and herbicidal activity of dibutyltin aryloxyacetate complexes[J]. J. Mol. Struct., 2025, 1320: 139476 doi: 10.1016/j.molstruc.2024.139476
-
[17]
JIANG W J, LUO Q, HUANG W, TAN Y X, PENG Y Y. Synthesis, anticancer activity, and mechanistic investigations of aryl-alkyl diorganotin arylformylhydrazone complexes[J]. J. Inorg. Biochem., 2025, 262: 112756 doi: 10.1016/j.jinorgbio.2024.112756
-
[18]
SHU S, ZHANG F X, TANG R H, YAN S Y, ZHU X M, SHENG L B, KUANG D Z, FENG Y L, YU J X, JIANG W J. Syntheses, structures and antitumor activities of tri(o-bromobenzyl)tin diethyldithiocarbamate and tri(m-fluorobenzyl)tin pyrrolidine dithiocarbamate[J]. Chin. J. Struct. Chem., 2020, 39(3): 459-466
-
[19]
LIU J, LIN Y C, LIU M, WANG S Q, LI Y X, LIU X C, TIAN L J. Synthesis, structural characterization and cytotoxic activity of triorganotin 5-(salicylideneamino)salicylates[J]. Appl. Organomet. Chem., 2019, 33(3): e4715 doi: 10.1002/aoc.4715
-
[20]
PAUL A, KHAN R A, SHAIK G M, SHAIK J P, RAI A K, DA SILVA M F C G, POMBEIRO A J L. Anthracene appended organotin(Ⅳ) compounds: Synthesis, structure elucidation and their cytotoxicity against A549 and RBL cancer cell lines[J]. Appl. Organomet. Chem., 2023, 37(3): e7232
-
[21]
GUL R, MUHAMMAD N, SIRAJUDDIN M, NOOR A, TUMANOV N, WOUTERS J, CHAFIK A, SOLAK K, MAVI A, SHUJAH S, ALI S, KHAN G S, QAYYUM S. Design, physicochemical confirmation, single crystal structures as well as exploration of antibacterial and anticancer potential of organotin(Ⅳ) carboxylates[J]. J. Mol. Struct., 2024, 1300: 137306 doi: 10.1016/j.molstruc.2023.137306
-
[22]
SHI H L, MA J W, LI Q L, DU X M, MENG Z X, RU J, MA C L. Four organotin(Ⅳ) complexes derived from 2, 6-difluoro-3-(propylsulfona-mido)benzoic acid: Synthesis, structure, in vitro cytostatic activity and antifungal activity evaluation[J]. Inorg. Chim. Acta, 2023, 551: 121485 doi: 10.1016/j.ica.2023.121485
-
[23]
张复兴, 邝代治, 王剑秋, 冯泳兰, 许志锋, 陈志敏, 曾荣英. 环状二聚三(邻甲基苄基)氢氧化锡的合成、结构及量子化学研究[J]. 有机化学, 2008, 28(8): 1457-1461ZHANG F X, KUANG D Z, WANG J Q, FENG Y L, XU Z F, CHEN Z M, ZENG R Y. Synthesis, crystal structure and quantum chemistry of the ring-form dimer tris(o-methylbenzyl)tin hydroxide[J]. Chin. J. Org. Chem., 2008, 28(8): 1457-1461
-
[24]
CHANDRASEKHAR V, THIRUMOORTHI R. Facile, ambient temperature, double Sn—C bond cleavage: Synthesis, structure, and electrochemistry of organotin and organotellurium ferrocene carboxylates[J]. Eur. J. Inorg. Chem., 2008(29): 4578-4585
-
[25]
ZHANG J H, ZHANG R F, MA C L, WANG D Q, WANG H Z. New organotin carboxylates derived from 6-chloro-3-pyridineacetic acid exhibiting discrete molecular, drum-like, linear polymeric and ladder structures constructed from dimeric tetraorganodistannoxane units[J]. Polyhedron, 2011, 30: 624-631 doi: 10.1016/j.poly.2010.11.035
-
[26]
AIRAPETYAN D V, PETROSYAN V S, GRUENER S V, ZAITSEV K V, ARKHIPOV D E, KORLYUKOV A A. Disproportionation reactions within the series of coordinated monoorganostannanes[J]. J. Organomet. Chem., 2013, 747: 241-248 doi: 10.1016/j.jorganchem.2013.07.005
-
[27]
IQBAL M, ALI S, MUHAMMAD N, PARVEZ M, LANGER P, VILLINGER A. Synthesis, characterization, crystal structures and electrochemical studies of organotin(Ⅳ) carboxylates[J]. J. Organomet. Chem., 2013, 723: 214-223 doi: 10.1016/j.jorganchem.2012.10.006
-
[28]
何丽芳, 唐文杰, 罗尧泽, 梁明勝, 唐建新, 吴雨萱, 张复兴, 朱小明. 基于2,2′-联吡啶-6,6′-二甲酸构筑的两个二烃基锡配合物的合成、结构及抗癌活性[J]. 无机化学学报, 2025, 41(8): 1601-1609HE L F, TANG W J, LUO Y Z, LIANG M S, TANG J X, WU Y X, ZHANG F X, ZHU X M. Synthesis, structure, and anticancer activity of two dialkyltin complexes constructed based on 2,2′-bipyridin-6,6′-dicarboxylic acid[J]. Chinese J. Inorg. Chem., 2025, 41(8): 1601-1609
-
[29]
SALAS J M, QUIRÓS M, ROMERO M A, SÁNCHEZ M P, SALAS M A. Metal complexes of theophylline-7-acetic acid. Crystal structure of a nickel(Ⅱ) compound containing noncoordinated theophylline-7-acetate ion[J]. Polyhedron, 1995, 14(5): 611-616 doi: 10.1016/0277-5387(94)00279-N
-
[30]
VIJAYANTHIMALA R, UDUPA M R. Complexes of palladium(Ⅱ) with theophylline-7-acetic acid[J]. Inorg. Chim. Acta, 1990, 175: 163-170 doi: 10.1016/S0020-1693(00)84823-9
-
[1]
-
Table 1. Crystallographic data and refinement parameters of the complex
Parameter [Sn(C6H11)3(tp)(H2O)] Empirical formula C27H44N4O5Sn Formula weight 623.35 Crystal system Orthorhombic Space group Iba2 a / nm 1.954 11(10) b / nm 1.702 11(9) c / nm 1.754 22(9) V / nm3 5.834 7(5) Z 8 Dc / (Mg·m-3) 1.419 μ(Mo Kα) / cm-1 9.17 F(000) 2 592 θ range for data collection / (°) 1.587-29.090 Index range -19 ≤ h ≤ 26,
-23 ≤ k ≤ 22,
-22 ≤ l ≤ 24Reflection collected 18 987 Reflection collected, unique 7 010 (Rint=0.033 2) Goodness-of-fit on F 2 1.025 Final R indices R1, wR2 [I > 2σ(I)] 0.031 9, 0.063 6 R indices R1, wR2 (all data) 0.049 1, 0.071 1 Largest diff. peak and hole / (e·nm-3) 614 and -346 Table 2. Selected bond distances (nm) and selected bond angles (°) of the title complex
Sn1—O3 0.213 1(3) Sn—C16 0.216 0(4) Sn1—O5 0.259 8(3) Sn1—C2 0.215 8(5) Sn1—C1 0.217 2(6) O3—Sn1—C2 91.72(19) C22—Sn1—C10 113.55(18) C22—Sn1—O5 82.99(17) O3—Sn1—C16 100.04(14) C16—Sn1—C10 126.8(2) C16—Sn1—O5 79.16(14) C22—Sn1—C16 114.83(19) O3—Sn1—O5 173.65(14) C10—Sn1—O5 86.15(17) O3—Sn1—C10 99.2(2) Table 3. Calculated some frontier molecular orbitals composition of the complex at the B3LYP/LANL2DZ level
ε / a.u. Sn O1 C1 L C2 C3 O2 H 158 -0.298 93 18.004 16 23.815 09 0.416 21 1.880 56 31.111 49 18.967 60 0.609 72 5.607 34 159 -0.282 94 1.727 00 41.748 25 1.286 45 52.236 33 1.529 51 0.530 76 0.409 20 1.824 77 160 -0.274 13 0.183 39 6.210 45 0.274 62 91.249 12 0.314 41 0.134 85 0.029 10 1.863 13 161 -0.271 55 1.567 41 82.126 74 2.184 50 4.520 87 7.452 52 3.317 19 0.005 92 1.004 55 162HO -0.222 71 4.157 04 1.460 73 0.388 42 89.181 90 3.090 14 0.051 13 0.020 44 2.030 33 163LU 0.236 16 0.153 17 1.646 15 1.079 03 91.609 41 0.108 92 0.018 57 0.005 94 0.649 58 164 0.321 81 53.155 39 2.408 11 0.284 80 0.719 14 28.141 96 5.905 84 6.643 74 3.014 85 165 0.330 03 0.299 42 0.089 97 0.165 88 98.279 37 0.265 69 0.099 08 0.014 06 0.948 18 166 0.344 15 0.472 36 0.196 20 0.908 53 98.452 24 0.284 39 0.051 23 0.000 68 0.535 92 167 0.359 22 3.120 34 40.046 67 47.584 89 48.463 97 2.425 80 0.417 19 0.007 57 5.516 55 Table 4. IC50 of the complex and paclitaxel on tumor cells in vitro
Compound IC50 / (μmol·L-1) AGS MOLT4 MDA-MB-321 [Sn(C6H11)3(tp)(H2O)] 2.6 1.433 7.061 Paclitaxel 0.894 4.587 0.004 -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 0
- 文章访问数: 356
- HTML全文浏览量: 38

 下载:
下载:






 下载:
下载:

