

含吡啶-2,6-二羧酸的钴(Ⅱ)配合物的合成、表征、抗肿瘤活性和理论计算
English
Synthesis, Characterization, Antitumor Activity, and Theoretical Calculations of Co(Ⅱ) Complex Based on Pyridine-2, 6-dicarboxylic Acid
-
Key words:
- cobalt(Ⅱ) complex
- / crystal structure
- / antitumor activity
- / DFT calculation
-
0. Introduction
Nowadays, cancers have become the most serious threat to human′s health worldwide[1]. It is reported that annual cancer cases will rise to 26 million within the next twenty years[2]. Unfortunately, there is no effective therapy for curing most of tumors up to now, and thus, it is still greatly desired to explore novel chemotherapeutic agents for oncotherapy[3-4]. Consequently, plentiful efforts have been devoted to exploring new antitumor drugs during the past decades. Among them, metallotherapeutics have attracted great attention since the discovery of cisplatin more than five decades ago[5-7]. Such metal-base drugs can prevent cancer cell division and then result in cancer cell apoptosis by inducing DNA damage and disrupting DNA repair process[8-9].
It is known that cisplatin is one of the most commonly used drugs in clinical treatment of numerous forms of human cancers, such as testicular and ovarian cancers, neck and head tumors[10-12]. Despite the great utility of cisplatin, its therapeutic value is also accompanied by serious side effects[13] and the increasing drug resistance[14]. This has created a continuous momentum in developing platinum - based drugs[15-16], and thus a few drugs, such as carboplatin, oxaliplatin, nedaplatin, lobaplatin, and heptaplatin have been successfully employed for clinical use in the past several decades. Such great successful clinical applications of platinum-based complexes as well as the accompanying toxicity and resistance have stimulated more research interests to search for other alternative transition metal (TM) coordination complexes in the area of metallotherapeutics.
In recent years, a great amount of Co-based compounds with antitumor activities have been achieved by employing different kinds of ligands[17-21]. To explore novel Co-based antitumor compounds, an interesting multidentate ligand, pyridine-2, 6-dicarboxylic acid (H2pdc) attracted our great interest because that it has a rigid 120° angle between the central pyridine ring and two carboxylate groups, and therefore could potentially provide various coordination modes to form both discrete and consecutive metal complexes[22]. Up to now, a large number of coordination compounds with intriguing topological structures have been constructed by using this ligand[23-29]. Motivated by the above information, a novel Co(Ⅱ)-based coordination compound [Co (Hpdc)(bpy)Cl]·C2H5OH (bpy=2, 2′-bipyridine) has been synthesized in this work. This complex was characterized by experimental analyses and theoretical computations. Theoretical results indicate that this complex is a good candidate for antitumor agents, and therefore, its inhibition against chronic myelocytic leukemia (K562) and esophageal carcinoma (OE-19) cells was determined by using thiazolyl blue tetrazolium bromide (MTT) assay. Our results reveal that this complex possesses certain antiproliferation effects on both of K562 and OE-19 cells with low IC50 values of (0.22±0.05) μg·mL-1 and (0.82±0.16) μg·mL-1, respectively.
1. Experimental and theoretical methods
1.1 Materials and physical measurements
All chemicals were purchased commercially and used without further purification. The K562 and OE-19 cells were obtained from Shanghai Institute of Materia Medica, Chinese Academy of Sciences. Elemental analyses (C, H, N) were carried out with a Perkin-Elmer 240 elemental analyzer. Infrared spectrum was recorded in a range of 400~4 000 cm-1 on a Perkin-Elmer Spectrum-2000 FT-IR spectrometer using KBr pellets.
1.2 Experimental synthesis
A mixture of H2pdc (2.05 mmol, 0.342 g) and 2, 2′bipyridine (1.20 mmol, 0.294 g) was dissolved in deionized water (10 mL). After stirring for 15 minutes, 1.20 mmol CoCl2·6H2O (0.286 g) was added into the above solution along with adding 5 mL absolute ethyl alcohol. Then, the mixture was continuously stirred at room temperature for 3 h, resulting in pellucid light green solution. After being filtrated and the filtration standing at room temperature for one week, reddish brown columnar crystals of [Co(Hpdc) (bpy)Cl]·C2H5OH were obtained (Yield: 55% based on Co). Anal. Calcd. for C19H18ClCoN3O5(%): C, 49.32; H, 3.92; N, 9.08. Found (%): C, 49.23; H, 3.99; N, 9.16. IR (KBr pellet, cm-1): 3 590~2 815 (br), 2 360 (w), 1 619 (vs), 1 430 (m), 993 (w), 734 (w).
1.3 Crystal structural determination
X-ray data of Co(Hpdc)(bpy)Cl·C2H5OH was collected on a Rigaku Saturn 724 CCD diffractometer equipped with a graphite-monochromator Mo Kα radiation (λ=0.071 073 nm) using an ω scan mode at 293(2) K. Absorption corrections were applied using multiscan methods. Data reduction was performed using the Rigaku Americas Corporation′s Crystalclear program[30]. This structure was solved by direct methods and refined by full-matrix least-squares using the SHELXS-97[31] and SHELXL-2018/1[32], respectively. All non-hydrogen atoms were refined anisotropically. Hydrogen atoms were located geometrically and treated as riding atoms with a common fixed isotropic thermal parameter.
CCDC: 1994088.
1.4 Computational details
The geometric structure of [Co(Hpdc)(bpy)Cl] with all real frequencies was optimized by using the PBE0 method in conjunction with the LANL2DZ basis set for Co atom and 6-31+G(d, p) basis set for the other atoms. The doublet structure with the lowest spin state is found to be the most stable molecular configuration of [Co(Hpdc) (bpy)Cl] by considering different spin states in structural optimization. The electrostatic potential (ESP), nature population analysis (NPA) charges, frontier molecular orbitals (FMO), and various electronic properties of [Co(Hpdc) (bpy)Cl] were obtained at the PBE0/6 -31++G(d, p) & LANL2DZ level based on the optimized configuration. Herein, chemical hardness (η), global electronegativity (χ), electrophilicity index (ω), chemical potential (CP) and softness (S) are defined as follows[33-35]:
$ {\chi = \frac{{\mid {\rm{VIE}} + {\rm{VEA}}\mid }}{2}} $ (1) $ {{\rm{CP}} = \frac{{|{\rm{VIE}} + {\rm{VEA}}|}}{2}} $ (2) $ {\eta = \frac{{{\rm{VIE}} - {\rm{VEA}}}}{2}} $ (3) $ {\omega = \frac{{{{({\rm{CP}})}^2}}}{{2\eta }}} $ (4) $ {S = \frac{1}{{2\eta }}} $ (5) To minimize the inaccuracy of the aforementioned parameters due to the self-interaction error (SIE) of density functional theory (DFT) methods[36-37], the vertical electronic affinity (VEA) and vertical ionization energy (VIE) were not directly approximated from the HOMO-LUMO energy (i. e., Koopman′s theorem). Instead, they are calculated by using the following equations[38]:
$ {\rm{VIE = }}{E_{{\rm{cation}}}}{\rm{ - }}{E_{{\rm{neutral}}}} $ (6) $ {\rm{VIE = }}{E_{{\rm{neutral}}}}{\rm{ - }}{E_{{\rm{anion}}}} $ (7) where Eneutral represents the electronic energy of the optimized neutral structure, while Ecation and Eanion are the cationic and anionic energies calculated at optimized neutral geometry, respectively.
All the above calculations were carried out by using the Gaussian 16 program package[39], in which the self-consistent reaction field (SCRF) calculations with the polarizable continuum model (PCM) [40] were performed to consider the solvent effect of water. Dimensional plots of molecular configurations and orbitals were generated with the GaussView program[41]. The electrostatic potential (ESP) was obtained by employing Multiwfn program package[42] in conjunction with the VMD software[43].
1.5 In vitro cytotoxicity
Cytotoxicity of this complex was assessed by MTT assay. Briefly, the K562 and OE-19 cells were cultivated into suspension cells in the 0.25% pancreatic protein. Then, the suspension liquid with 1×105 cells per milliliter was seeded into a 96-well culture plate with 100 μL per well. After incubation at 37 ℃ in a 5% (V/V) CO2 incubator for 24 h, the samples containing the synthesized complex of serial concentrations were added into the wells of the experimental groups (10 μL per well). DMSO without this complex was added into the well of the control group (10 μL per well). The cells were sequentially incubated for 72 h, followed by the addition of 20 μL MTT solution (5 mg·mL-1) to each well and further cultivation for 4 h. Afterwards, the supernatant with MTT was removed and followed by the addition of 150 μL DMSO to dissolve formazan dye for 10 min at room temperature along with slow oscillation. Finally, the optical density (OD) at 490 nm was read by an enzyme-linked immunosorbent assay (ELISA) reader and the inhibition rate (IR=(1-ODcomplex/ODblank)×100%) were calculated. The median inhibitory concentration (IC50) values were obtained from the results of quadruplicate determinations of at least three independent experiments. Herein, the samples containing the titled complex were obtained by dissolving the isolated crystals into the DMSO following the high temperature sterilization. Thereafter, these solutions were diluted by RPMI 1640 nutrient solution to the concentrations of 100, 50, 25, 12.5, 6.25, 3.13, 1.56, and 0.78 μg·mL-1.
2. Results and discussion
2.1 Crystal structure
The synthesized complex [Co(Hpdc) (bpy)Cl]·C2H5OH appears as a reddish brown columnar crystal. Herein, the crystal structure of this complex is shown in Fig. 1 in conjunction with the theoretically optimized molecular structure of [Co(Hpdc)(bpy)Cl], while the 3D network is given in Fig. S1 (Supporting information). Crystallographic data are collected in Table 1, and selected bond lengths and angles from experimental and theoretical characterization are listed in Table S1.
Figure 1
 Figure 1. Molecular structures of (a) [Co(Hpdc)(bpy)Cl]·C2H5OH obtained by experiment and (b) [Co(Hpdc)(bpy)Cl] in water from theoretical calculation with atom numbering
Figure 1. Molecular structures of (a) [Co(Hpdc)(bpy)Cl]·C2H5OH obtained by experiment and (b) [Co(Hpdc)(bpy)Cl] in water from theoretical calculation with atom numberingTable 1
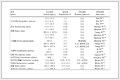 Table 1. Crystallographic data collection and structure refinement details of [Co(Hpdc)(bpy)Cl]·C2H5OH
Table 1. Crystallographic data collection and structure refinement details of [Co(Hpdc)(bpy)Cl]·C2H5OHEmpirical formula C19H18ClCoN3O5 Formula weight 462.74 Crystal system Monoclinic Space group P21/n a / nm 1.089 8(2) b / nm 1.712 5(3) c / nm 1.141 2(2) β / (°) 105.29(3) V / nm3 2.054 3(8) Z 4 μ/ mm-1 1.001 Dc / (g·cm-3) 1.496 θ range / (°) 3.01~27.48 Index ranges -14 ≤ h ≤ 14, -22 ≤ k ≤ 22, -13 ≤ l ≤ 14 Reflection collected 19 862 Unique reflection 4 696 Observed reflection [I>2σ(I)] 4 095 F(000) 948 Rint 0.024 7 Goodness of fit on F2 0.986 R1 [I > 2σ(I)]a 0.031 0 wR2 [I>2σ(I)]b 0.081 7 R1 (all data) 0.037 1 wR2 (all data) 0.085 5 (Δρ)max, (Δρ)min / (e·nm-3) 579, -315 a R1=∑||Fo|-|Fc||/∑|Fo|; b wR2={∑[w(Fo2-Fc2)2]/∑[w(Fo2)2]}1/2 As shown in Table 1, the crystal structure of [Co(Hpdc) (bpy)Cl]·C2H5OH belongs to P21/n space group of monoclinic system. As shown in Fig. 1a, its molecular structure in the crystal is composed of one Co2+, one 2, 6-Hpdc- anion, one 2, 2′-bipyridine, one Cl-, and one guest alcohol molecule. It is observed that the Co2+ ion in this complex is six-coordinated: two carboxylate oxygen atoms and one nitrogen atom from the pdc2- ligand, two nitrogen atoms from 2, 2′bipyridine, and one Cl- anion. As shown in Table S1, the bond lengths of Co—N1, Co—N2, Co—N3, Co— O1, Co—O3, and Co—Cl are 0.208 43(13), 0.212 80(14), 0.209 60(16), 0.218 12(13), 0.232 20(14), and 0.234 26(10) nm, respectively, which are consistent with those of previously reported Co-based complexes[44-45]. These subunits of [Co(Hpdc)(bpy)Cl]·C2H5OH are linked together by hydrogen bonds to form a stable 3D framework (Fig.S1), and the values of hydrogen bond lengths and angles for [Co(Hpdc) (bpy)Cl] ·C2H5OH are listed in Table S2.
It is known that the practical application of such complex usually needs to dissolve it into water. Thus, it is meaningful to further reveal the electronic structure and relevant properties of this novel complex dissolved in water. So, the isolated [Co(Hpdc)(bpy)Cl] motif from experimental crystal structure has been re-optimized at the PBE0/6-31+G(d, p) & LANL2DZ level by considering the water solvent. Herein, the ethanol molecule in this complex has been neglected because that ethanol is easily dissolved in water and then has little effect on the structure and properties of [Co(Hpdc) (bpy)Cl] in aqueous solution. After optimization, it is found that the calculated Co—N1 (0.195 6 nm), Co—N2 (0.198 0 nm), Co—N3 (0.200 0 nm), Co—O1 (0.215 9 nm), and Co—Cl (0.233 8 nm) bond lengths are slightly shorter than those obtained by the single crystal XRD experiment, while the bond length of Co—O3 (0.245 5 nm) is longer than that from the experiment. And the differences between the theoretical and the experimental values of ∠N—Co—N and ∠O—Co—O angles are smaller than 9°. Hence, the optimized Co(Hpdc) (bpy)Cl in aqueous solution exhibits a very similar geometric structure to that obtained from crystal characterization, indicating that the used theoretical method in this work is reliable. The slight inconsistencies in the bond lengths and angles are commonly found in previously reported literatures[44, 46-48], which can be attributed to the fact that the environment of crystal lattice in the experiment is different from the aqueous condition used in theoretical simulation.
2.2 Experimental and theoretical characterization
Firstly, the experimental IR spectrum of this studied complex has been shown in Fig. S2. Herein, the strong vibrations appearing around 1 620 and 1 400 cm-1 correspond to the asymmetric and symmetric stretching vibrations of the carboxylate group, respectively. The absorption bands in a range of 650~670 cm-1 and 720~750 cm-1 in the fingerprint region are attributed to Co—O and Co—N vibrations, respectively.
In view of the fact that the synthesized complex will become a monomer after being dissolved into water, the molecular model from theoretical optimization will be closer to its real state after entering cells. Therefore, a detailed study on the electronic structure of this complex was performed on the basis of the theoretically optimized [Co(Hpdc) (bpy)Cl]. The charge distribution, ESP, FMO, and various global reactivity parameters of [Co(Hpdc) (bpy)Cl] in aqueous solution will be discussed in the following sections.
To clearly reveal the distribution of charges in this complex, the calculated NPA charges of [Co(Hpdc) (bpy)Cl] are displayed in Fig.S3. It can be seen that the negative charges mainly distribute on the electronegative O, N, and Cl atoms in [Co(Hpdc)(bpy)Cl], while Co carries a positive charge of 0.580. In addition, the ESP picture of [Co(Hpdc) (bpy)Cl] is given in Fig. 2. As shown in Fig. 2, the positive electrostatic potential mainly locates on peripheral hydrogen atoms of Hpdc- and bpy ligands. It is found that the hydrogen of carboxyl in [Co(Hpdc)(bpy)Cl] has the largest positive potential of 317.02 kJ·mol-1, indicating that this hydrogen atom has a large tendency to form hydrogen bond with N or O atoms of DNA bases when it approaches DNA in cells. In another respect, the Cl atoms in [Co(Hpdc) (bpy)Cl] are surrounded by large negative potential, which is consistent with the NBO analysis. Also, the HOMO and LUMO orbitals of [Co(Hpdc)(bpy) Cl] are depicted in Fig. 3. It is observed that this complex has large HOMO-LUMO gaps of 4.30 and 4.40 eV for α and β electrons, indicating that it also has high chemical stability.
Figure 2
Figure 3
To further reveal the electronic properties of this complex, the global reactivity parameters, including VEA, VIE, η, χ, ω, CP, and S of [Co(Hpdc) (bpy)Cl] were also calculated and summarized in Table S3. These theoretical results may provide some references for experimentalists in the future. In addition, it is known that the molecular dipole moment has an obvious effect on the intermolecular interaction between drug molecules and drug targets[47]. As shown in Table S3, the dipole moment of [Co(Hpdc)(bpy)Cl] was found to be 7.17 a.u., indicating that this complex is soluble in polar solvent, such as water. Moreover, it is reported that the increase in polarizability of aromatic ring complexes can enhance the interaction between them and the DNA of tumor cells[24]. From Table S3, it is found that, with two ligands containing aromatic rings, the polarizability of [Co(Hpdc)(bpy)Cl] is as large as 376 a.u., suggesting that this complex is also likely to interact with DNA. These static electric properties again imply that this complex is a good candidate to serve as a new antitumor metallotherapeutic agent. Consequently, it is of great interest to evaluate the antitumor activity of this complex in experiment.
2.3 In vitro cytotoxicity
The growth inhibitory effect of [Co(Hpdc)(bpy)Cl]· C2H5OH on two kinds of human tumor cells, i.e., K562 and OE-19 cells in vitro were evaluated by MTT assay. The inhibition rates on K562 and OE-19 under different concentrations of this complex are listed in Table S4 and plotted in Fig. 4. As shown in Fig. 4, it is observed that the inhibition rates on K562 and OE-19 gradually increased along with the increasing concentrations of this complex. In particular, the estimated IC50 values of this complex for K562 and OE-19 cells were (0.22±0.05) μg·mL-1 and (0.82±0.16) μg·mL-1 (i.e., (0.48±0.11) μmol·L-1 and (1.77±0.35) μmol·L-1), respectively. These small IC50 values demonstrate that this complex is quite effective in restraining the growth of K562 and OE-19 cells. Besides, it is noted that the IC50 value for K562 was much smaller than that for OE19, indicating that this complex is more cytotoxic for K562 cells. More interestingly, the IC50 value ((0.22± 0.05) μg·mL-1) of [Co(Hpdc) (bpy)Cl] ·C2H5OH on K562 cell was less than those of 0.70~1.86 μg·mL-1 previously reported for cisplatin[49-52], which further verifies the inhibitory effect of this Co-based compound on K562 cells.
Figure 4
3. Conclusions
In summary, a Co(Ⅱ)-based coordination compound [Co(Hpdc) (bpy)Cl] ·C2H5OH has been synthesized and characterized by IR spectrum, elemental analysis, and single-crystal X-ray diffraction. Also, the electronic structure of this complex in water was also detailedly studied by using DFT calculations. Moreover, its antitumor activity was experimentally studied by MTT assay in K562 and OE-19 cancer cell lines. The experimental results show that this novel complex indeed exhibits a significant antitumor activity against K562 and OE-19 cells, which demonstrates its great potential of serving as a new antitumor agent. This work will not only provide a novel Co-based complex as potential antitumor agent, but also intrigue more interest in exploring the synthesis and application of new coordination complexes in biomedical field.
Supporting information is available at http://www.wjhxxb.cn
#共同第一作者。
-
-
[1]
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin D M, Forman D, Bray F. Int. J. Cancer, 2015, 136(5):E359-E386 doi: 10.1002/ijc.29210
-
[2]
Thun M J, DeLancey J O, Center M M, Jemal A, Ward E M. Carcinogenesis, 2009, 31(1):100-110
-
[3]
Tomasetti C, Li L, Vogelstein B. Science, 2017, 355(6331):1330-1334 doi: 10.1126/science.aaf9011
-
[4]
Livingston D M, Silver D P. Nature, 2008, 451(7182):1066-1067 doi: 10.1038/4511066a
-
[5]
Rosenberg B, Vancamp L, Krigas T. Nature, 1965, 205(4972):698-699 doi: 10.1038/205698a0
-
[6]
Rosenberg B, Vancamp L, Trosko J E, Mansour V H. Nature, 1969, 222(5191):385-386 doi: 10.1038/222385a0
-
[7]
Hurley, Laurence H. Nat. Rev. Cancer, 2002, 2(3):188-200 doi: 10.1038/nrc749
-
[8]
Hartinger C G, Dyson P J. Chem. Soc. Rev., 2009, 38(2):391-401 doi: 10.1039/B707077M
-
[9]
Thota S, Rodrigues D A, Crans D C, Barreiro E J. J. Med. Chem., 2018, 61(14):5805-5821 doi: 10.1021/acs.jmedchem.7b01689
-
[10]
Bakalova A, Varbanov H, Buyukliev R, Momekov G, Ferdinandov D, Konstantinov S, Ivanov D. Eur. J. Med. Chem., 2008, 43(5):958-965 doi: 10.1016/j.ejmech.2007.06.025
-
[11]
Galanski M, Arion V B, Jakupec M A, Keppler B K. Curr. Pharm. Des., 2003, 9(25):2078-2089 doi: 10.2174/1381612033454180
-
[12]
Tanaka T, Yukawa K, Umesaki N. Oncol. Rep., 2005, 14(5):1365-1369
-
[13]
Rabik C A, Dolan M E. Cancer Treat. Rev., 2007, 33(1):9-23 doi: 10.1016/j.ctrv.2006.09.006
-
[14]
Ohmichi M, Hayakawa J, Tasaka K, Kurachi H, Murata Y. Trends Pharmacol. Sci., 2005, 26(3):113-116 doi: 10.1016/j.tips.2005.01.002
-
[15]
Galanski M. Recent Patents Anti-Canc. Drug Discov., 2006, 1(2):285-295 doi: 10.2174/157489206777442287
-
[16]
Shaili E. Sci. Prog., 2014, 97(1):20-40 doi: 10.3184/003685014X13904811808460
-
[17]
陈战芬, 马艺丹, 华罗光, 张健.无机化学学报, 2014, 30(7):1525-1534 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20140706&flag=1CHEN Z F, MA Y D, HUA L G, ZHANG J. Chinese J. Inorg. Chem., 2014, 30(7):1525-1534 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20140706&flag=1
-
[18]
Pires B M, Giacomin L C, Castro F A V, Amanda D S C, Pereira M D, Bortoluzzi A J, Faria R B, Scarpellini M. J. Inorg. Biochem., 2016, 157:104-113 doi: 10.1016/j.jinorgbio.2016.01.024
-
[19]
解庆范, 郭妙莉, 陈延民.无机化学学报, 2018, 34(2):309-316 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20180213&flag=1XIE Q F, GUO M L, CHEN Y M. Chinese J. Inorg. Chem., 2018, 34(2):309-316 http://www.wjhxxb.cn/wjhxxbcn/ch/reader/view_abstract.aspx?file_no=20180213&flag=1
-
[20]
Li J, Zhang J, Zhang Q, Wang Y, Bai Z, Zhao Q, He D, Wang Z, Zhang J, Chen Y. Bioorg. Med. Chem., 2019, 27(20):115071 doi: 10.1016/j.bmc.2019.115071
-
[21]
Khan H Y, Ansari M O, Shadab G G H A, Tabassum S, Arjmand F. Bioorg. Chem., 2019, 88(2019):102963
-
[22]
Chuasaard T, Panyarat K, Rodlamul P, Chainok K, Yimklan S, Rujiwatra A. Cryst. Growth Des., 2017, 17(3):1045-1054 doi: 10.1021/acs.cgd.6b01389
-
[23]
Xu J, Su W, Hong M. Cryst. Growth Des., 2011, 11(1):337-346 doi: 10.1021/cg101343k
-
[24]
Ghosh S K, Ribas J, Bharadwaj P K. CrystEngComm, 2004, 6(45):250-256 doi: 10.1039/B407571D
-
[25]
Bordbar M, Tabatabaee M, Alizadeh-Nouqi M, Mehri-Lighvan Z, Khavasi H R, YeganehFaal A, Fallahian F, Dolati M. J. Iran. Chem. Soc., 2016, 13(6):1125-1132 doi: 10.1007/s13738-016-0826-x
-
[26]
Ghosh S K, Bharadwaj P K. Inorg. Chem., 2004, 43(7):2293-2298 doi: 10.1021/ic034982v
-
[27]
Ghosh S K, Bharadwaj P K. Inorg. Chem., 2005, 44(9):3156-3161 doi: 10.1021/ic048159q
-
[28]
Derikvand Z, Dorosti N, Hassanzadeh F, Shokrollahi A, Mohammadpour Z, Azadbakht A. Polyhedron, 2012, 43(1):140-152 doi: 10.1016/j.poly.2012.06.026
-
[29]
Deng D, Liu P, Fu W, Li L, Yang F, Ji B. Inorg. Chim. Acta, 2010, 363(5):891-898 doi: 10.1016/j.ica.2009.12.044
-
[30]
CrystalClear 1.40, Rigaku Americas Corp, The Woodlands, TX, 2008.
-
[31]
Sheldrick G M. SHELXL-97, Program for the Solution of Crystal Structures, University of Göttingen, Germany, 1997.
-
[32]
Sheldrick G M. SHELXL-2018/1, Program for the Refinement of Crystal Structures, University of Göttingen, Germany, 2018.
-
[33]
Hutama A S, Huang H, Kurniawan Y S. Spectrochim. Acta. A Mol. Biomol. Spectrosc., 2019, 221:117152
-
[34]
Parr R G, Donnelly R A, Levy M, Palke W E. J. Chem. Phys., 1978, 68(8):3801-3807 doi: 10.1063/1.436185
-
[35]
Parr R G, Szentpaly L V, Liu S. J. Am. Chem. Soc., 1999, 121(9):1922-1924
-
[36]
Cohen A J, Mori-Sánchez P, Yang W. Science, 2008, 321(5890):792-794 doi: 10.1007%2F128_2011_195
-
[37]
Mori-Sanchez P, Cohen A J, Yang W. J. Chem. Phys., 2006, 125(20):2604-308
-
[38]
Lu C, Kuang X Y, Lu Z W, Mao A J, Ma Y M. J. Phys. Chem. A, 2011, 115(33):9273-9281 https://pubmed.ncbi.nlm.nih.gov/21780834/
-
[39]
Frisch M J, Trucks G W, Schlegel H B, Scuseria G E, Robb M A, Cheeseman J R, Scalmani G, Barone V, Petersson G A, Nakatsuji H, Li X, Caricato M, Marenich A V, Bloino J, Janesko B G, Gomperts R, Mennucci B, Hratchian H P, Ortiz J V, Izmaylov A F, Sonnenberg J L, Williams-Young D, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski V G, Rega N, Gao J, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery JrJA, Peralta J E, Ogliaro F, Bearpark M J, Heyd J J, Brothers E N, Kudin K N, Staroverov V N, Keith T A, Kobayashi R, Normand J, Raghavachari K, Rendell A P, Burant J C, Iyengar S S, Tomasi J, Cossi M, Millam J M, Klene M, Adamo C, Cammi R, Ochterski J W, Martin R L, Morokuma K, Farkas O, Foresman J B, Fox D J. Gaussian 16, Revision A.03, Gaussian, Inc., Wallingford, 2016.
-
[40]
Scalmani G, Frisch M J. J. Chem. Phys., 2010, 132(11):114110
-
[41]
Dennington R, Keith T, Millam J. GaussView, Ver. 6, KS, USA: Semichem Inc, Shawnee Mission, 2016.
-
[42]
Lu T, Chen F J. Comput. Chem., 2012, 33(5):580-592
-
[43]
Humphrey W, Dalke A, Schulten K. J. Mol. Graphics, 1996, 14(1):33-38
-
[44]
Stamatatos T C, Pringouri K V, Raptopoulou C P, Vicente R, Psycharis V, Escuer A, Perlepes S P. Inorg. Chem. Commun., 2006, 9(12):1178-1182
-
[45]
Marques L F, Marinho M V, Speziali N L, Visentin L D C, Machado F C. Inorg. Chim. Acta, 2011, 365(1):454-457 https://www.sciencedirect.com/science/article/abs/pii/S0020169310005384
-
[46]
Ghasemi K, Rezvani A R, Shokrollahi A, Moghimi A, Gavahi S, García-Granda S, Mendoza-Meroño R. C. R. Chim., 2014, 17(12):1221-1229
-
[47]
Gao H. Spectrochim. Acta Part A Mol. Biomol. Spectrosc., 2011, 79(3):687-693
-
[48]
Štarha P, Marek J, Trávníček Z. Polyhedron, 2012, 33(1):404-409
-
[49]
Krstic N M, Matic I Z, Juranic Z D, Novakovic I T, Sladic D M. J. Steroid. Biochem. Mol. Biol., 2014, 143:365-375
-
[50]
Ohe Y, Nakagawa K, Fujiwara Y, Sasaki Y, Minato K, Bungo M. Cancer Res., 1989, 49(15):4098-4102
-
[51]
Han Q B, Li R T, Zhang J X, Sun H D. Helv. Chim. Acta, 2004, 87(5):1119-1124 doi: 10.1002/chin.200444202
-
[52]
Su W C, Chang S L, Chen T Y, Chen J S, Tsao C J. Jpn. J. Clin. Oncol., 2000, 30(12):562-567
-
[1]
-
Table 1. Crystallographic data collection and structure refinement details of [Co(Hpdc)(bpy)Cl]·C2H5OH
Empirical formula C19H18ClCoN3O5 Formula weight 462.74 Crystal system Monoclinic Space group P21/n a / nm 1.089 8(2) b / nm 1.712 5(3) c / nm 1.141 2(2) β / (°) 105.29(3) V / nm3 2.054 3(8) Z 4 μ/ mm-1 1.001 Dc / (g·cm-3) 1.496 θ range / (°) 3.01~27.48 Index ranges -14 ≤ h ≤ 14, -22 ≤ k ≤ 22, -13 ≤ l ≤ 14 Reflection collected 19 862 Unique reflection 4 696 Observed reflection [I>2σ(I)] 4 095 F(000) 948 Rint 0.024 7 Goodness of fit on F2 0.986 R1 [I > 2σ(I)]a 0.031 0 wR2 [I>2σ(I)]b 0.081 7 R1 (all data) 0.037 1 wR2 (all data) 0.085 5 (Δρ)max, (Δρ)min / (e·nm-3) 579, -315 a R1=∑||Fo|-|Fc||/∑|Fo|; b wR2={∑[w(Fo2-Fc2)2]/∑[w(Fo2)2]}1/2 -

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 5
- 文章访问数: 1603
- HTML全文浏览量: 193

 下载:
下载:



 下载:
下载:

