 图式 1
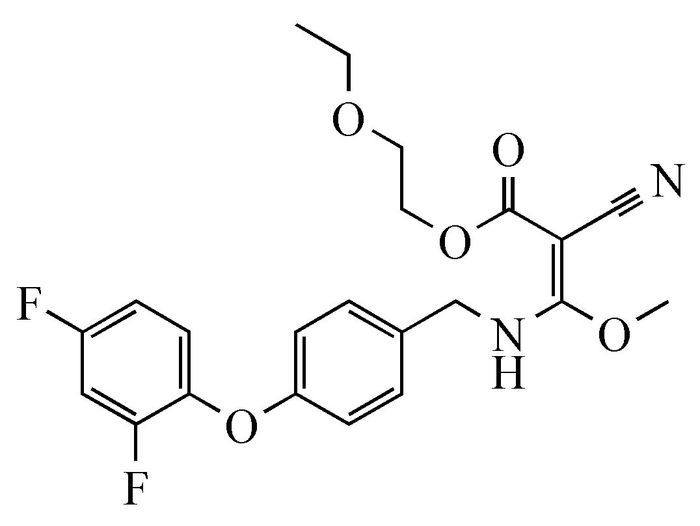
The structure of MAE
Scheme1.
The structure of MAE
图式 1
The structure of MAE
Scheme1.
The structure of MAE

(E)-2-氰基-3-[4-(2, 4-二氟苯氧基)-苄氨基]-3-甲氧基-丙烯酸2-乙氧基乙酯对超螺旋质粒DNA的特异性切割
English
Specific Cleavage of Supercoiled Plasmid DNA by (E)-2-Cyano-3-[4-(2, 4-difluorophenoxy) benzylamino]-3-methoxyacrylic acid 2-Ethoxyethyl Ester
-
Key words:
- DNA cleavage
- / Acrylic ester
- / Spectroscopy
- / DNA sequencing
-
DNA carrying hereditary information is one of essential macromolecules for all known living organisms, and the cleavage of DNA is of paramount importance in molecular biology, gene engineering and biochemistry[1~3]. Usually, the DNA cleavage can be done with restriction enzymes. However, a variety of organic small molecule compounds with entirely different structures were found to cleave DNA efficiently at specific sequence or site over the last decade, which have been developed as artificial nucleases or chemical nucleases[4~6]. Compared with endonucleases, artificial nucleases have telling advantages in terms of design flexibility and ease of manufacture, which makes the active study on the interaction between chemical nuclease and DNA of current general interest and importance.
With advances in biotechnology and medicinal chemistry, most of today's synthetic DNA cleavage agents are metal complexes.[7] Such metal ions in metallonucleases can promote DNA scission and enhance the efficiency or specificity of nucleic acid cleavage. However, the metal-ion-promoted DNA cleavage, more often than not, involves free-radical and oxidative mechanism which cannot simulate the chemistry of natural restriction endonucleases to yield 5'-OPO3 and 3'-OH ends. In addition, dopant ions such as cadmium, copper are harmful even in high dilution[8]. Therefore, the development of organic compounds is an emerging area of interest for DNA hydrolysis.
It has been reported that cyanoacrylates have high biological activities and have been widely used as herbicides, fungicides, anticancer drugs, etc[9, 10]. A cyanoacrylic ester, (E)-2-cyano-3-[4-(2, 4-difluorophenoxy)benzylamino]-3-methoxy-acrylic acid 2-ethoxy-ethyl ester, named as MAE (Scheme 1), is designed and synthesized to study its structure-dependent herbicidal activity[11]. Unexpectedly, MAE possesses DNA cleavage activity. In this work, the interaction between MAE and DNA is discussed, the DNA cleavage activity of cyanoacrylic esters is investigated, and a possible cleavage mechanism is proposed simultaneously.
1 Experimental
1.1 Materials and reagents
MAE is synthesized according to the method reported in the literature[11]. 1H NMR (400 MHz, CDCl3) δ: 9.66 (br, 1H, NH), 7.18 (d, 2H, J=8.8Hz, Ar-H), 7.04~7.10 (m, 1H, Ar-H), 6.84~6.98 (m, 4H, Ar-H), 4.44 (d, 2H, J=6.0Hz, CH2NH), 4.28 (t, 2H, J=5.2Hz, COOCH2CH2), 4.23 (s, 3H, CH3O), 3.68 (t, 2H, J=5.2Hz, COOCH2CH2), 3.56 (q, 2H, J=7.2Hz, CH2CH3), 1.20 (t, 3H, J=7.2Hz, CH2CH3). Anal. Calcd for C22H22F2N2O5: C, 61.11; H, 5.13; N, 6.48. Found: C, 60.76; H, 5.10; N, 6.62.
Original 100 mmol/L of MAE solution was prepared with DMSO as dissolvent, and diluted with respective amount of Tri-HCl buffer (pH 7.4) to various proper concentrations for all experiments. The plasmid pUC19 DNA was amplified and extracted from E. coli Top10 according to the published method[12]. Plasmid DNA was dissolved into 1 mmol/L Tris-HCl buffer (pH 7.4) and stored at -20℃. The DNA concentration was determined by measuring the absorbance at 260 nm. The original DNA solution got a 260/280 ratio of around 1.8, which indicated that the DNA was sufficiently free of protein.
1.2 Absorption spectroscopy
Ultraviolet-visible (UV-Vis) spectra were performed using a double beam UV-Vis 2550 spectrophotometer (Shimadzu, Japan) with a pair of quartz cuvettes (1cm light path), and Tris-HCl buffer, containing an appropriate amount of DMSO, was used as reference solution. The absorption spectra of a series of solutions, successive addition of DNA into the 5 mL volumetric flask containing a constant concentration of the compound MAE, were measured by scanning the UV spectrum from 200 to 400 nm.
1.3 Fluorescence spectroscopy
The fluorescence spectra were recorded on a Shimadzu RF-5301PC fluorescence spectrophotometer (Japan) in a 1cm quartz cell with slit widths of 5nm. The excitation and emission wavelengths were 475nm and 500~620nm with sampling point every 0.5nm, respectively. Fluorescence measurements were carried out with DNA concentration fixed at 5.78×10-6 mol/L, and the compound concentration varied from 33 to 67 μmol/L. Ethidium bromide (EB) was used as a fluorescent tag in this study.
1.4 Agarose gel electrophoresis
A 10 μL reaction solution containing 5 μg DNA and 0.02 μmol MAE was incubated for a certain period of time at 37℃, and then the cleavage reaction was quenched by adding 2 μL loading buffer (30 mmol/L EDTA, 36% glycerol, 0.05% xylene cyanol FF, 0.05% bromophenol blue). Gel electrophoresis was performed at 100 V for 30 min in TAE buffer (40 mmol/L Tris-acetate, 1 mmol/L EDTA) using 0.7% agarose gel. After stained with EB (1.0 mg/mL), DNA fragments in the gel were graphed by an electrophoresis imaging analysis system (BIO-Best 200E, SIM, USA) and semiquantitatively analyzed using the Bindscan software.
1.5 DNA sequence analysis
The double-stranded scission fragment was purified with agarose gel DNA purification kit (Dingguo, China), and then ligated using the Rapid DNA Ligation Kit (TakaRa, Japan). The ligation product of ds-DNA was transformed into E. coli competent cells Top10, and then analyzed by digestion with three restriction endonuclease including NdeⅠ, PstⅠ, and AseⅠ. For DNA sequencing, a sequence primer was designed with sequence as 5′-CTA GAG TAA GTA GTT CGC CAG-3′. At last, a quick alignment was carried out between cleavage fragment sequence and pUC19 original series using NCBI BLAST.
2 Results and Discussion
2.1 Absorption spectra
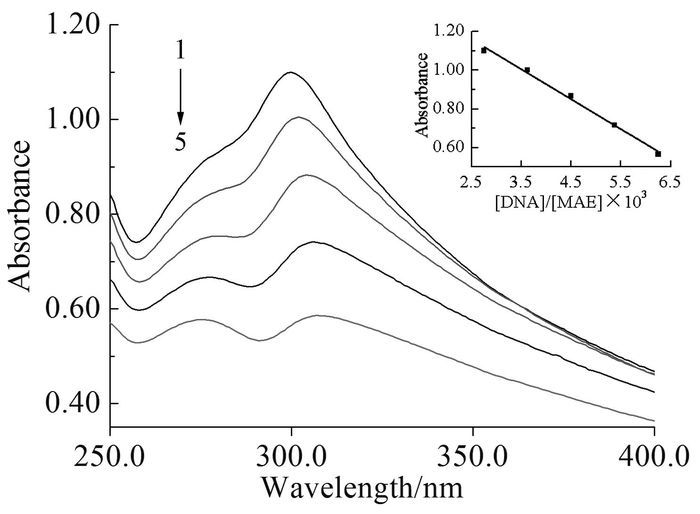
UV-Vis is an efficient technique to investigate the mode of interaction between exogenous substances and biological macromolecules, such as nucleic acid[13]. In general, red shift and hypochromism in the absorption spectra are found when compound binds to DNA helix[14]. The absorption spectra of the MAE-DNA system are measured with variable concentrations of DNA. As shown in Fig. 1, a bathochromic shift (8nm) and a hypochromism effect (47%) are observed with the concentration of MAE fixed at 80 μmol/L and DNA concentration increasing from 0.22 to 0.50 μmol/L. In addition, the plot of absorption with the ratio of DNA to MAE gives a good linear relation (r2=0.9915). With regard to these spectral results, it can be reasonably proposed that MAE binds to base pairs of DNA.
2.2 Fluorescence spectra
Generally, previous studies have shown that there is no fluorescence of DNA, and EB can intercalate double stranded DNA, which could intensify the fluorescence by almost 20-fold. Moreover, the fluorescent intensity of DNA/EB complex can be gradually decreased by the addition of the compound that can compete with EB in intercalating to DNA[15]. As shown in Fig. 2, there is an appreciable reduction in the emission intensity with the increasing of cyanoacrylic ester concentration, which is a proof that the compound MAE inserts into base pairs of DNA. The kinetic of fluorescence quenching is further illustrated using the Stem-Volmer equation[16]. With regard to Fig. 2, it is found that a plot of F0/F versus the concentration of MAE yields a straight line (r2=0.9986), and that the overall quenching constant (Ksv) is 1.12×104 L/mol, which means that the fluorophore is statically quenched by forming a stable complex between macromolecule DNA and quencher MAE, and is in accordance with the UV experimental results.
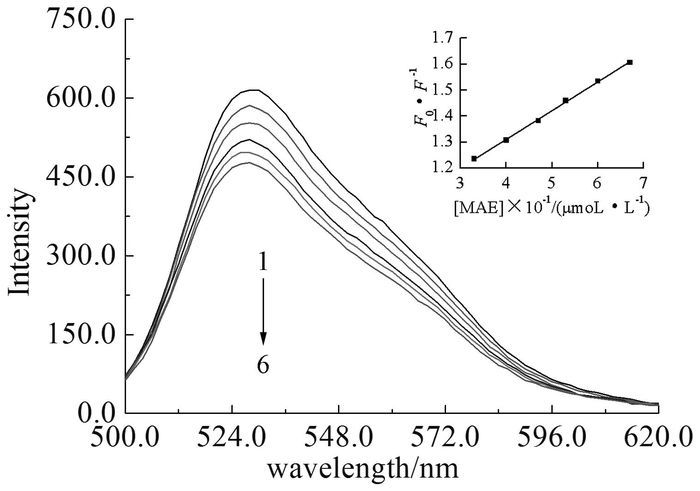 图2
Fluorescence spectra of DNA/EB system in the presence of increasing amount of MAE (Insert: Stern-Volmer plot of the interaction between DNA and compound MAE)
Figure2.
Fluorescence spectra of DNA/EB system in the presence of increasing amount of MAE (Insert: Stern-Volmer plot of the interaction between DNA and compound MAE)
图2
Fluorescence spectra of DNA/EB system in the presence of increasing amount of MAE (Insert: Stern-Volmer plot of the interaction between DNA and compound MAE)
Figure2.
Fluorescence spectra of DNA/EB system in the presence of increasing amount of MAE (Insert: Stern-Volmer plot of the interaction between DNA and compound MAE)
2.3 Agarose gel electrophoresis
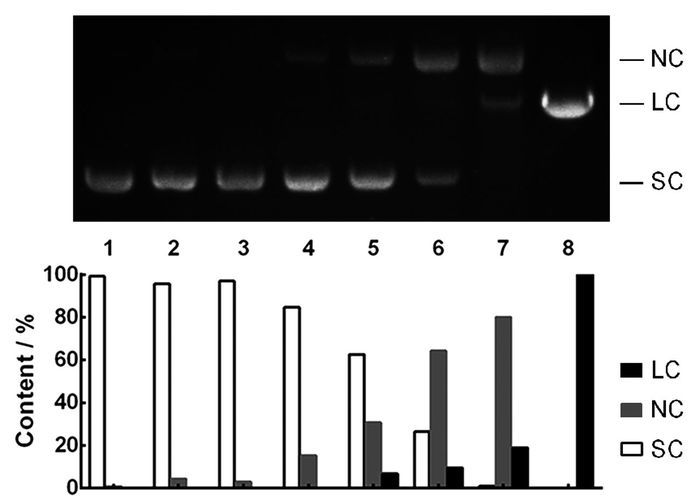
The synthetic product's DNA cleavage activity is studied by agarose gel electrophoresis using supercoiled pUC19 plasmid DNA as a substrate. Typically, there are three types of the plasmid DNA[17], namely supercoiled (SC), linear (LC) and nicked circles (NC), and DNA migration rate varies in the order: SC>LC>NC. The time-dependent cleavage reaction and gel electrophoresis are performed according to "Material and Methods", and the experimental results are shown in Fig. 3 with the image to the up showing a gel stained with EB and relative contents of three DNA forms listing below. The conversion of plasmid DNA from supercoiled to nicked circles is observed after incubation for 6 h (lane 2), and the linear form of DNA appeared in 24 h (lane 5). At last, there is only linear pUC19 DNA at 72 h (lane 8), suggesting that MAE exhibits a remarkable DNA cleavage activity.
2.4 DNA sequencing
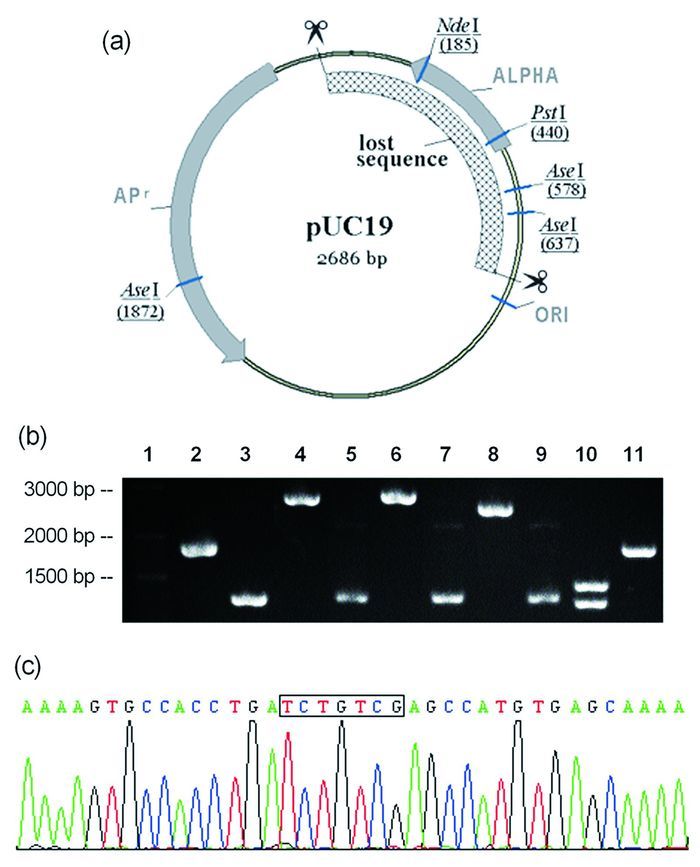
On the basis of the spectroscopic and electrophoretic observations, we focus our attention on whether the DNA cleavage of cyanoacrylate MAE is region-or sequence-specific mode. Digestion and sequence analysis are performed. Briefly, the double stranded scission fragment is recovered and recombined into a supercoiled DNA, named as pUC19-MAE. The restriction patterns of the original plasmid pUC19 and the recombinant vector pUC19-MAE are shown in Fig. 4. Digestion of the pUC19 DNA with Pst Ⅰ (lane 3) or Nde Ⅰ (lane 5) results in a fragment about 2700 bp, whereas no distinct digested products is observed following restriction digestion of pUC19-MAE with Pst Ⅰ (lane 4) or Nde Ⅰ (lane 6). The results are further verified by double-digestion of Pst Ⅰ and Nde Ⅰ. Digestion with Ase Ⅰ, pUC19 yields two clear bands as expected (lane 10) compared with only one fragment produced from pUC19-MAE (lane 11). Restriction analysis results show that a fragment containing the multiple cloning site in frame with the lacZ gene may be cut off. Upon DNA sequencing and BLAST sequence analysis, it is found that a sequence of 874 nucleotides is deleted from plasmid pUC19, suggesting that the compound MAE may cleave nucleic acid efficiently at position 2616 and 806 of plasmid DNA.
In addition, the plasmid DNA cleavage with cyanoacrylate MAE will be expected via the hydrolytic mechanism, not the oxidative mode in this work. One reason for this is that the in situ formation of reactive oxygen radical is essential for the oxidative DNA cleavage. Otherwise the reaction would be interrupted. Unfortunately, DMSO used as solvent to prepare original solution of MAE is a standard free radical scavenger, so we had to exclude the possibility of oxidative mode. The other is that the DNA fragment cleaved by the compound MAE can be rejoined orbicularly by T4 DNA ligase, which is in accord with the typical hydrolytic cleavage.
3 Conclusions
The DNA cleavage properties of cyanoacrylic ester MAE were investigated by absorption, fluorescence, gel electrophoresis and DNA sequencing. Considering the above UV absorption and fluorescence test results, cyanoacrylate MAE can intercalate between bases of plasmid DNA to form a stable complex. In gel electrophoresis, the amount of supercoiled type DNA is on the decline whereas those of nicked and linear DNA increase progressively with the increase in cleavage time. In addition, supercoiled DNA is converted to nick style and then to liner form, no direct double-strand DNA cleavage is shown.
On the basis of above evidences, the cleavage mechanism of the plasmid pUC19 DNA by artificial nuclease MAE can be generally proposed as a three-step process: DNA intercalation with MAE, site-specific single-stranded cleavage and then double-stranded hydrolytic scission, which will give a fresh impetus to site-specific DNA cleavage by artificial nuclease.
-
-
[1]
E Ma, L B Harrington, M R O'Connell et al. Mol. Cell, 2015, 60(3):398~407. doi: 10.1016/j.molcel.2015.10.030
-
[2]
S Gambino, B Mousley, L Cathcart et al. Anal. Biochem., 2016, 495:3~5. doi: 10.1016/j.ab.2015.11.013
-
[3]
A R Lambert, J P Hallinan, B W Shen et al. Structure, 2016, 24(6):862~873. doi: 10.1016/j.str.2016.03.024
-
[4]
T K N Luong, I Govaerts, J Robben et al. Chem. Commun., 2017, 53(3):617~620. doi: 10.1039/C6CC08555E
-
[5]
F Z Li, F M Feng, J Y Wu et al. Prog. React. Kinet. Mec., 2016, 41(1):39~47. doi: 10.3184/146867816X14490554565537
-
[6]
S Thota, S Vallala, R Yerra et al. Int. J. Biol. Macromol., 2016, 82:663~670. doi: 10.1016/j.ijbiomac.2015.09.045
-
[7]
Q Cheng, F Zhang, H Zhou et al. J. Coord. Chem., 2015, 68(11):1997~2005. doi: 10.1080/00958972.2015.1032272
-
[8]
C X Yuan, Y B Wu, Y B Wei et al. Chin. J. Chem., 2007, 25:1267~1272. doi: 10.1002/(ISSN)1614-7065
-
[9]
刘阳, 刘明, 陈明桂等.有机化学, 2017, 37:403~410. http://kns.cnki.net/KCMS/detail/detail.aspx?filename=yjhu201702012&dbname=CJFD&dbcode=CJFQ
-
[10]
S K Bagherpour Doun, S E Alavi, M K Moftakhari Esfahani et al. Tumour Biology, 2014, 35(8):7491~7497. doi: 10.1007/s13277-014-1996-8
-
[11]
胡方中, 李春晖, 潘俊辛等. CN:101967115B, 2013.
-
[12]
K J Humphreys, A E Johnson, K D Karlin et al. J. Biol. Inorg. Chem., 2002, 7:835~842. doi: 10.1007/s00775-002-0369-8
-
[13]
Y Zhao, K J Du, S Q Gao et al. J. Inorg. Biochem., 2016, 156:113~121. doi: 10.1016/j.jinorgbio.2016.01.004
-
[14]
J Lin, R T Liu, C Z Gao et al. Spectrochim. Acta A Mol. Biomol. Spectrosc., 2012, 97: 532~535.
-
[15]
N P Andreou, K Dafnopoulos, C Tortopidis et al. J Photochem. Photobiol. B, 2016, 158:30~38. doi: 10.1016/j.jphotobiol.2016.02.017
-
[16]
S Nahid, H Leila. Spectrochim. Acta. A Mol. Biomol. Spectrosc., 2012, 97:406~410.
-
[17]
A Arbuse, M Font, M A Martinez et al. Inorg. Chem., 2009, 48(23):11098~11107. doi: 10.1021/ic901488j
-
[1]
-
Figure 4 The cleavage sits of the plasmid pUC19
(a):pUC19 plasmid map. Every restriction enzyme site is shown by an underline. Cleavage sites of DNA by MAE are shown by two clipper symbols. (b): Agarose gel electrophoresis of pUC19-MAE DNA incised by different enzymes. lane 1: Normal RunTM prestained 250bp-Ⅱ DNA ladder; lane 2, 3: pUC19 DNA and pUC19-MAE control; lane 4, 6, 8 10: pUC19 digested with Pst Ⅰ, Nde Ⅰ, Pst Ⅰ/Nde Ⅰ, Ase Ⅰ, respectively; lane 5, 7, 9 11: pUC19MAE digested with Pst Ⅰ, Nde Ⅰ, Pst Ⅰ/Nde Ⅰ, Ase Ⅰ, respectively. (c): Partial sequencing result of pUC19-MAE. The recombination sequences are boxed
-

 扫一扫看文章
扫一扫看文章
计量
- PDF下载量: 0
- 文章访问数: 0
- HTML全文浏览量: 0


 下载:
下载:




 下载:
下载:

